Bayesian Networks can be seen as Markov Random Fields (MRFs), let’s see how:
 If there is a single edge, then it’s easy since the function can be interpreted as .
If there is a single edge, then it’s easy since the function can be interpreted as .
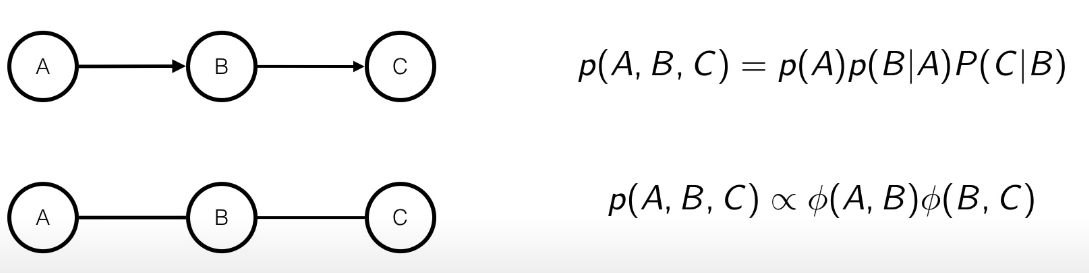 In the case there are more edges, then the parametrization is not unique, but it’s quite easy too, since we can interpret the potential function as:
In the case there are more edges, then the parametrization is not unique, but it’s quite easy too, since we can interpret the potential function as:
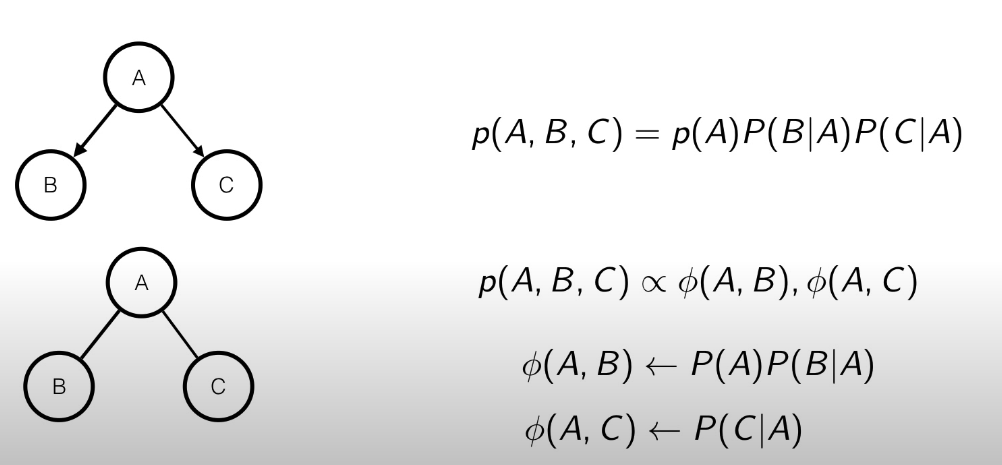 In this case, as we can see that we can model the potential functions as shown in the figure. Even here the parametrization is not unique, since we could bring the inside the .
In this case, as we can see that we can model the potential functions as shown in the figure. Even here the parametrization is not unique, since we could bring the inside the .
The problem
The three cases we’ve seen before are quite easy to represent using a MRF, but what if we have something like this?
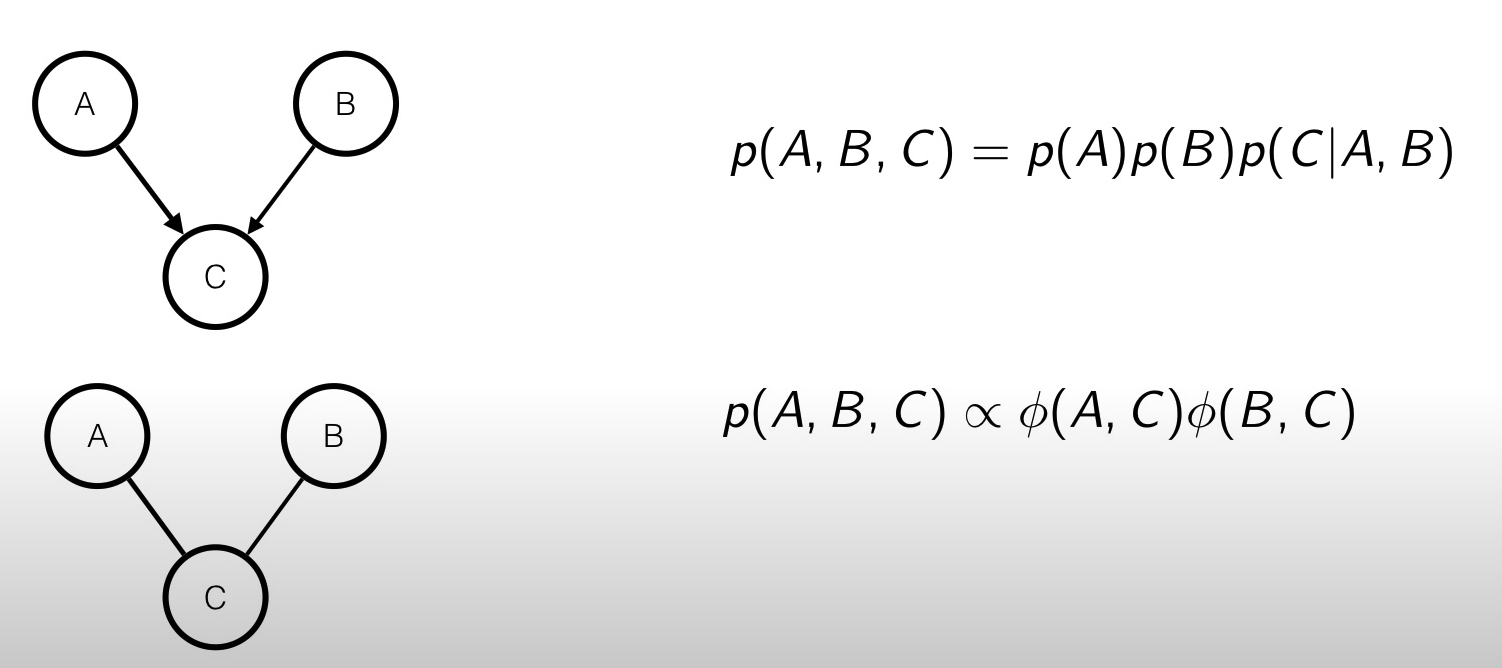 We can see that and are dependent given in the Bayesian Network, but not in the MRF.
We can see that and are dependent given in the Bayesian Network, but not in the MRF.
The solution is to moralize the parents, meaning that we have to connect the parents:
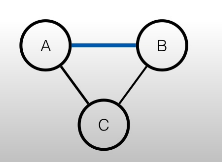 In order to have
In order to have